Tutorials
1. Quick Search for the interaction network of individual drug
The users could input a drug generic name or synonym (such as Acetylsalicylic acid, ABC) in the search bar of the home page to query the whole interaction network of the input drug. If the typed terms could match with multiple drug entries in the database, a list of suitable suggestions will be provided to users, with hyperlinks to corresponding full pages. Sometimes the number of matched entries is zero, possibly because of error input or unrecorded drugs. Note that not all the approved drugs are included in the current release, and the interaction networks of some drugs have never been identified. The query can be submitted by clicking the searching icon.
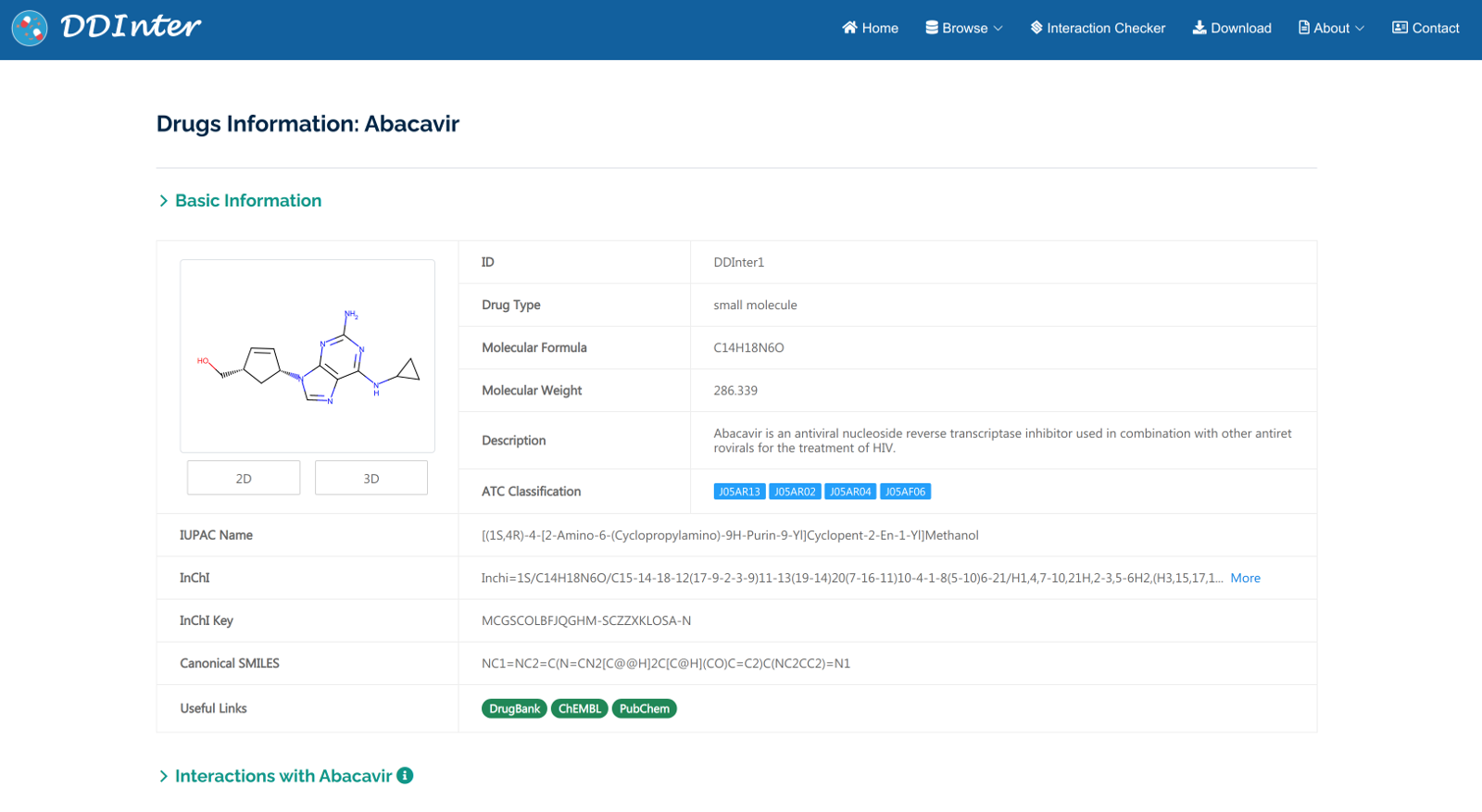
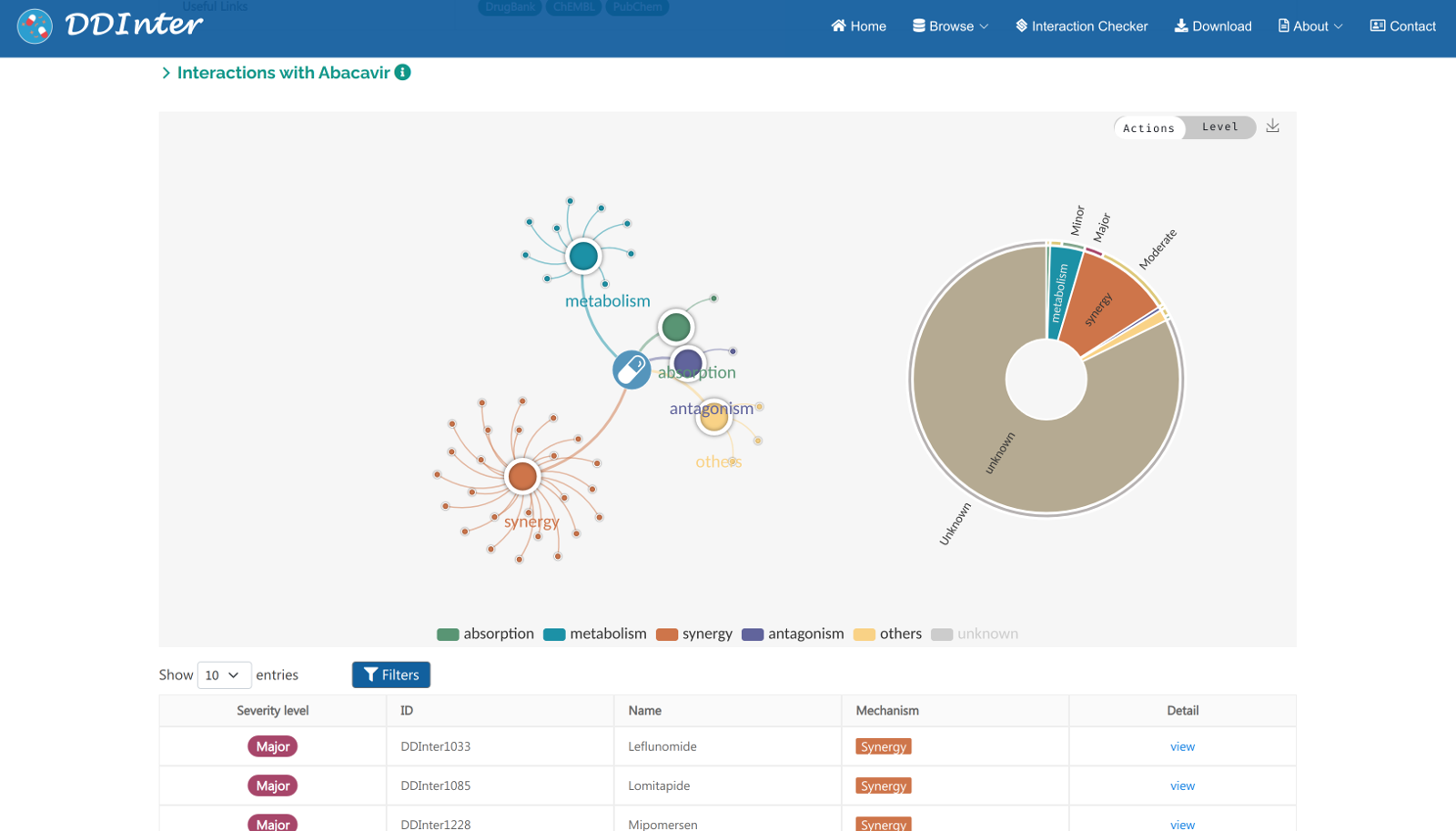
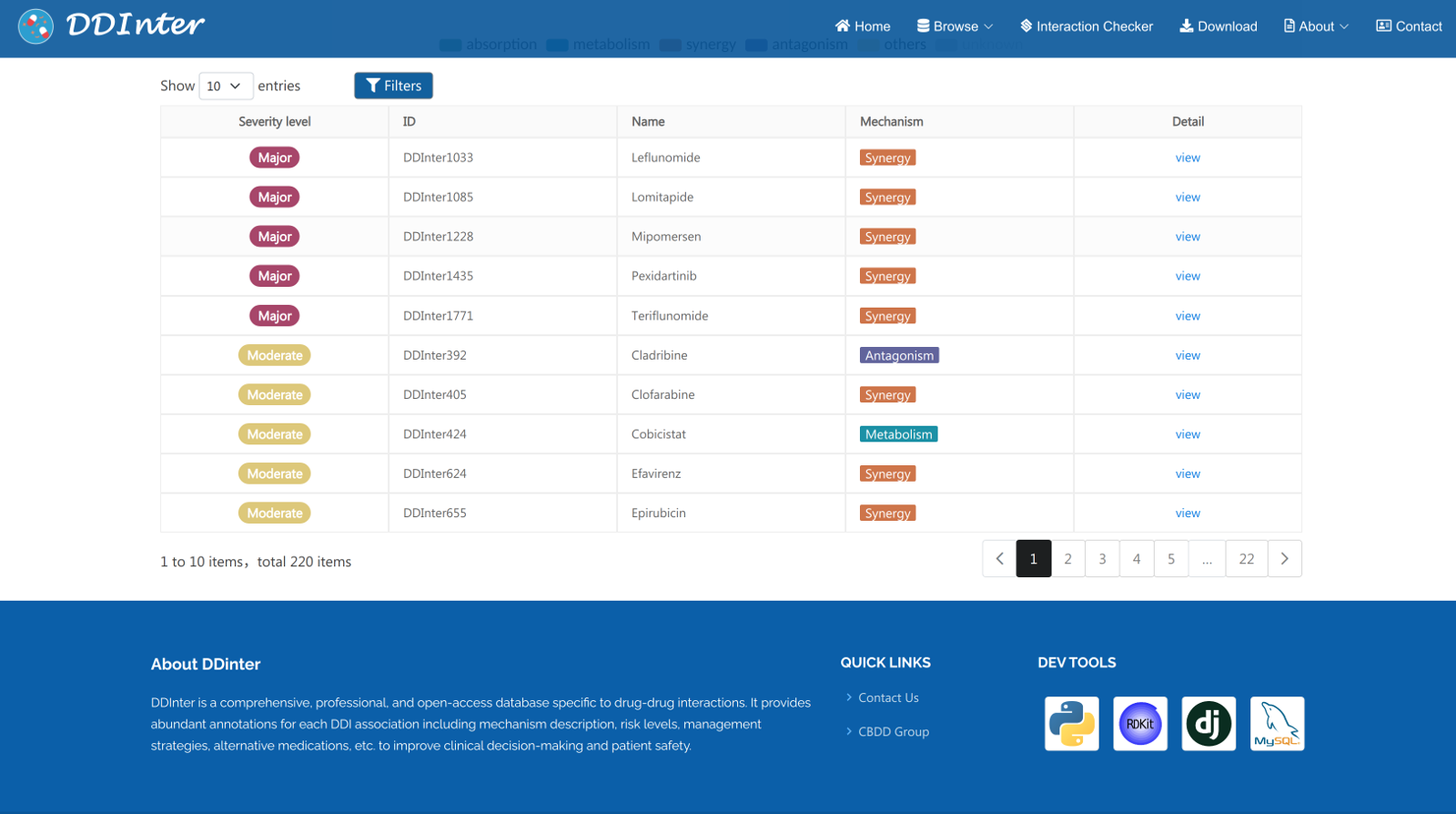
The resulting webpage includes basic information of the queried drug, two interactive charts displaying the distributions of DDI association, and a detailed table listing all the DDIs. (1) Basic information section displays drug ID, drug type, molecular formula, molecular weight, pharmacological description, ATC codes, chemical identifiers, and some useful external links. (2) A relation chart and a sunburst chart are used to visualize all the interacted terms dynamically, including severity level, interaction mechanism, name of interacted drugs, and proportion of each type. (3) The Interaction table displays severity level, ID of the interacted drug, name of interacted drug, mechanism, and the “View’ button linking to the detailed interaction page.
By clicking the “view” button in the interaction table, the detailed interaction information page of drug pairs will be displayed. The ‘Interaction’ field shows expanded descriptions mechanisms, like pharmacologic conflicts, synergetic toxicity, metabolic enzymes competition, etc. The ‘Management’ field displays strategies for managing potential side effects, including avoiding combinations, monitoring potential toxicity, adjusting the dosage of drugs, changing time of administration, etc. If available, the source from which the information was extracted is also provided to allow users to trace back to the original documents. The alternatives of each drug in DDI associations are based on the third level (pharmacological subgroup) of ATC codes. And the alternative medications have no interactions with the other drug. The metabolism chart is used to show potential associations mediated by metabolism enzymes. The metabolism data is predicted using ADMETlab 2.0, a web-accessible tool released by our group, and probability results higher than 0.7 (incline to be substrate or inhibitors) are marked with balloon icons. Users can browse concrete predicted values, switch the chart types and scaling, and download the full chart by using the operation broad in the upper-right corner.
2. Check for potential DDI risks in prescriptions
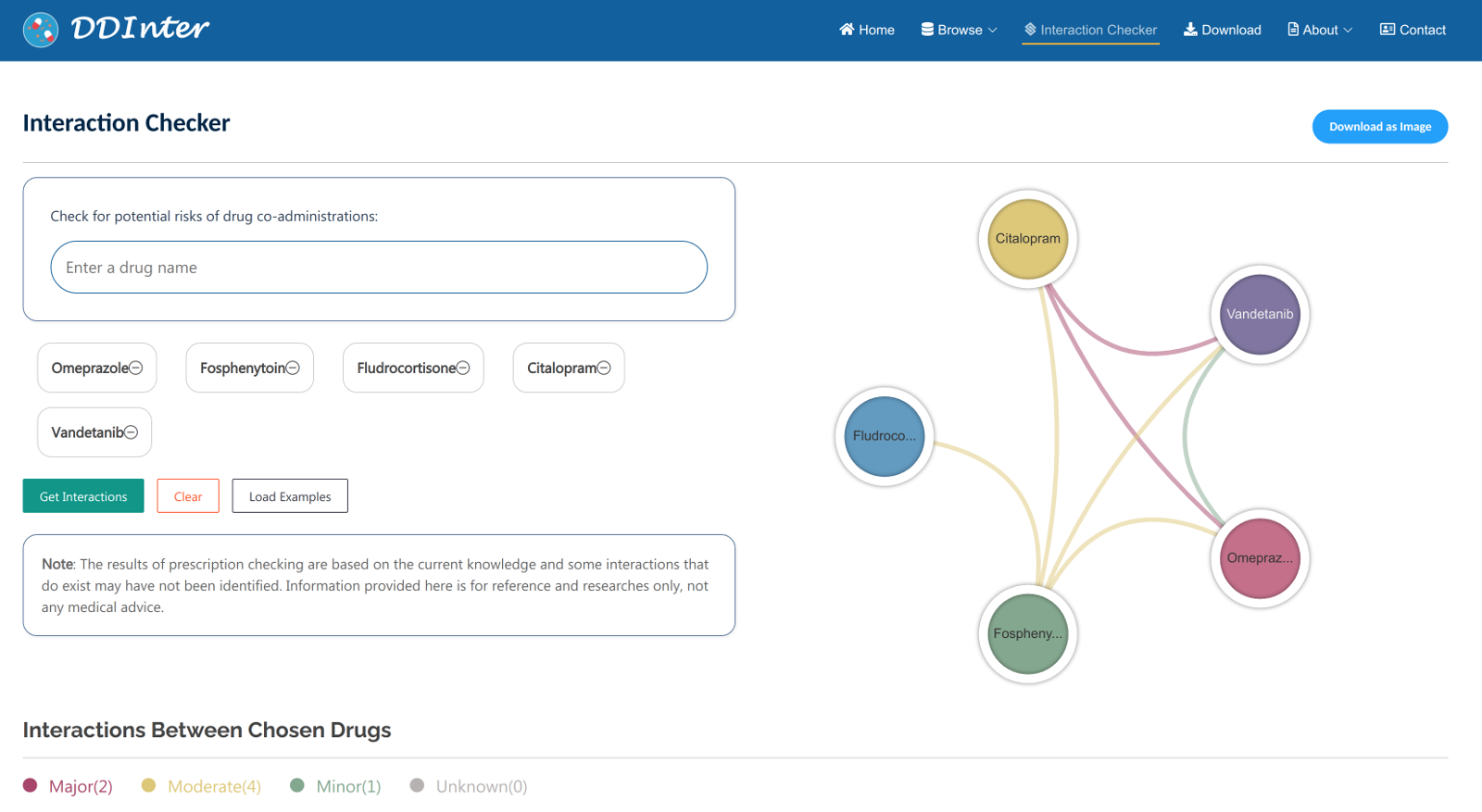
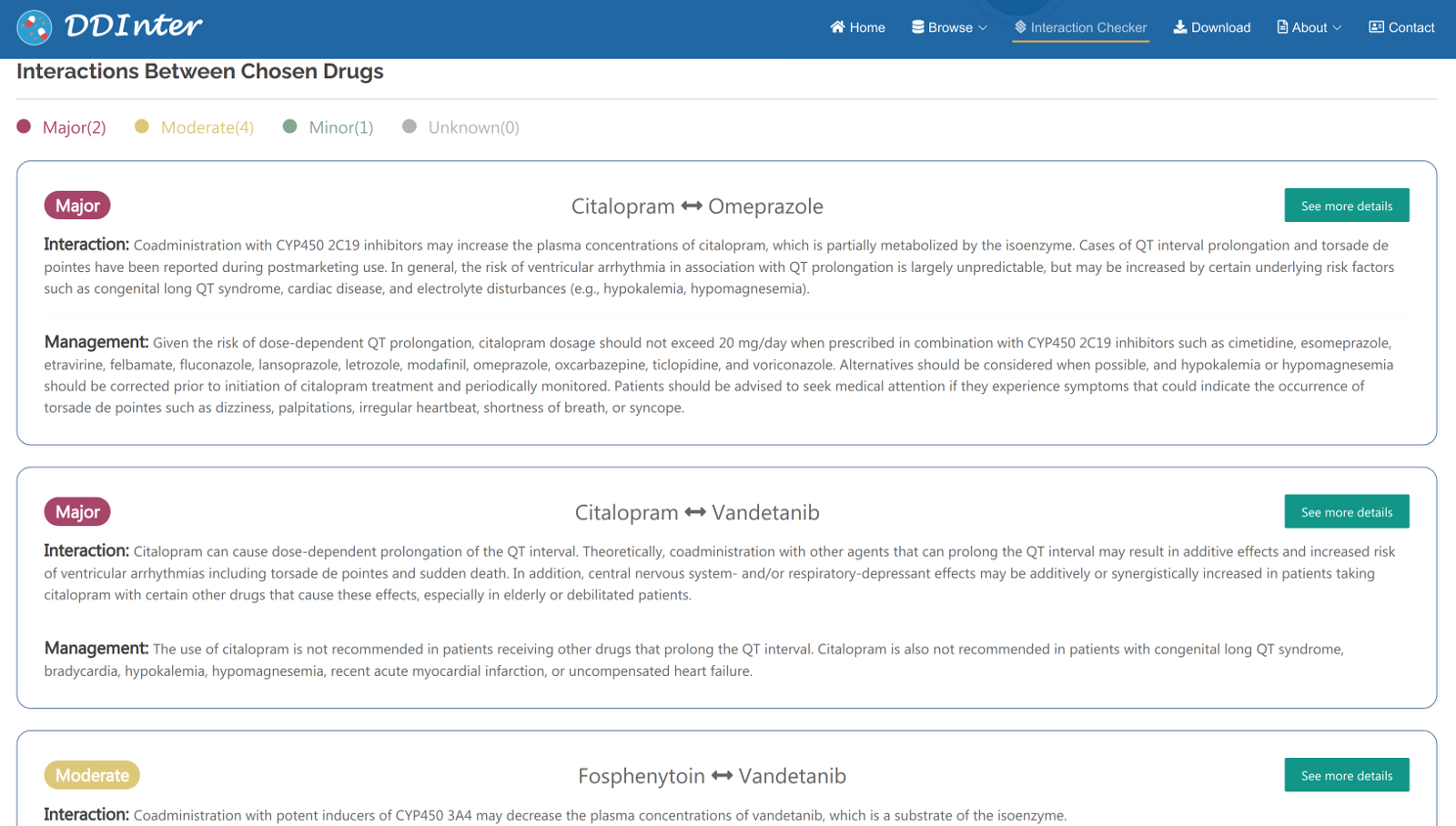
In the interaction checker module, users could check for potential risks of co-administration by entering a list of drugs (more than 2 drugs and up to 5drugs). First, users should enter a drug name and select the best match from the list of suggestions, and repeat the process to add multiple drugs. By clicking “Get Interactions” button, the information of the interacted drug pairs will be shown in multiple cards below. Users could see the risk level of specific drug pairs, mechanisms, and strategies for managing side effects. A relation chart of the chosen drugs is used to assist the users to understand the associations. By clicking “See more details” button, the interaction information page of drug pairs will be displayed. Click the “cross sign” in the chosen drugs bar to refresh the page for a new check.
3. Browse DDInter
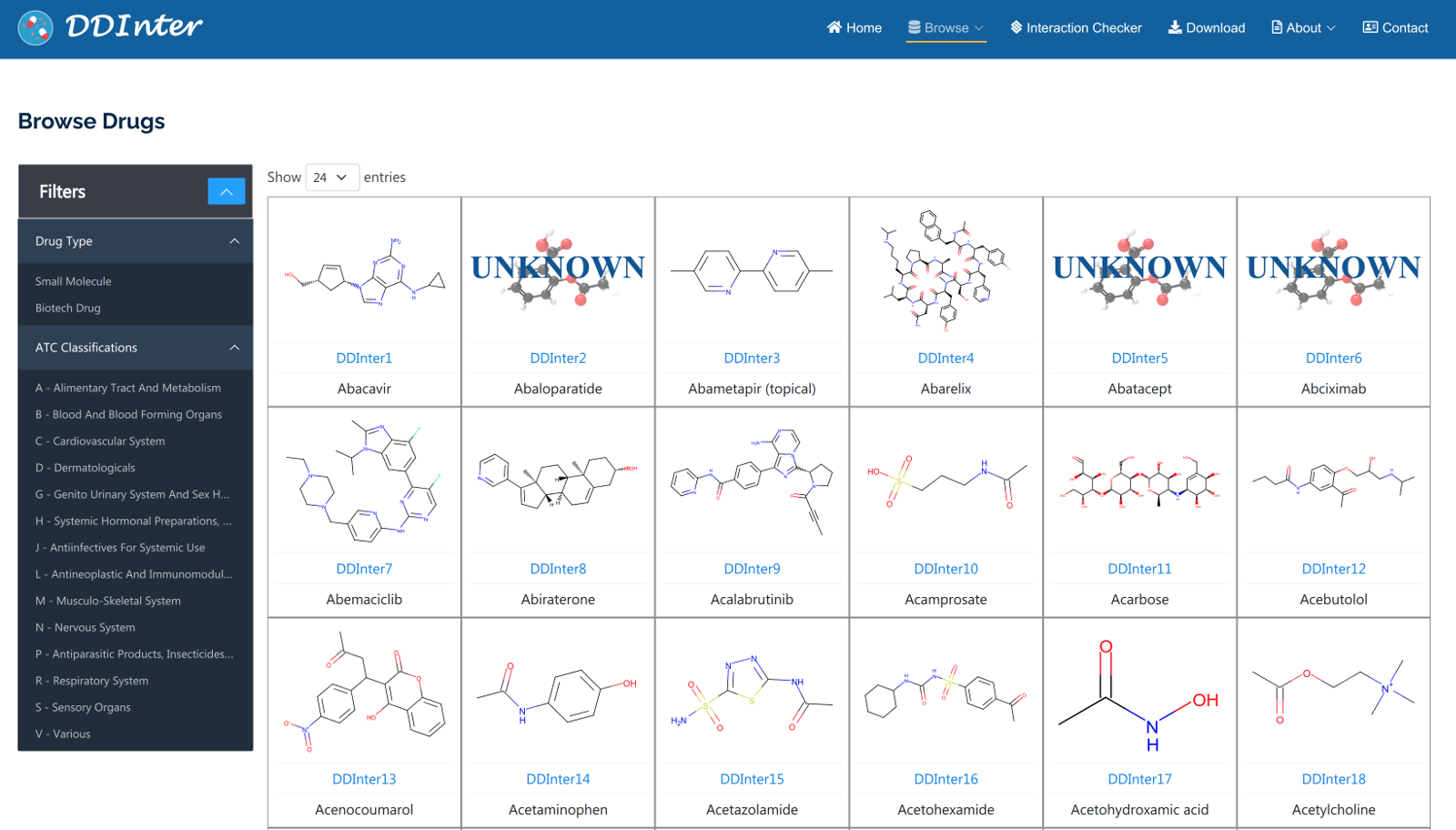
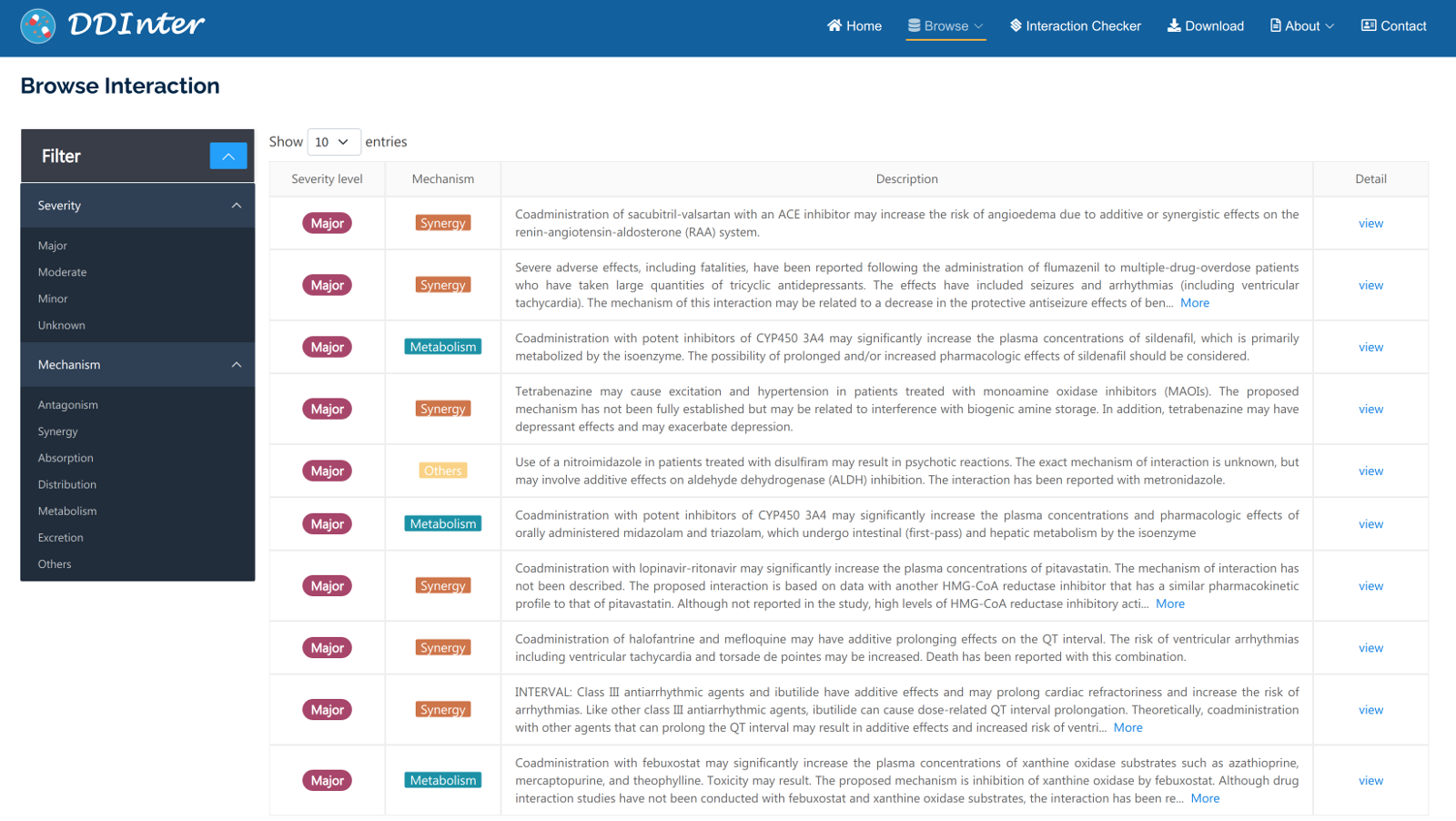
Two types of entries, drugs and interactions, could be browsed. All the drugs entities are assigned with unique DDInter identifiers and shown in the form of molecular structures. The interactive filter located on the left side of the page allows users to explore a subset of the original data, such as small-molecule drugs, biotech drugs, and drugs targeting different therapeutic areas. Clicking on a specific drug ID will open the drug information page that displays affluent interaction information as well as chemical and pharmacological information. For interaction browsing, severity level and mechanism filters are both provided to guide the users find entries of interest. These interaction entities are shown in the tabular format with basic information. When clicking on the ‘view’ button of specific interaction, a list of drug-drug pairs corresponding to this entry would be accessed, where users could obtain concrete interaction content from the hyperlinks provided. By default, drug entries and interaction entries are presented in order of identifier and risk level, respectively.
4. Downloads
All the DDI associations can be downloaded from the website without login or registration.